Text(1.2, 1.1, labels = "Log10 normalized counts", pos = 4) Loop to draw rectangles within the color key pal2 <- c(pal1, rev(pal)) Generate the y coordinates for rectangles within the color key rect_series = seq(0.3, 1, length.out = max(network_df$means) + 1) Text(l, l, labels=rownames(netm), pos = 3, cex = 0.7, font = 2) 5, lor = network_df$cols, vertex.size = 5, lor = This is the part that i want to draw in Cytoscape plot(network_graph, rescale = F, layout=l*1.0, vertex.label = NA, 
Provide breaks for the values of the nodes, so that they fall in a different color shade pal <- colorRampPalette(c("red", "slateblue"))(6)īks <- pheatmap:::generate_breaks(values, length(pal), center = F)Ĭols <- pheatmap:::scale_colours(values, col=pal, breaks=bks, na_col = "grey") Normalization of coordinates, so we know how large the figure is in base graphics coordinates l <- norm_coords(l, ymin=-1, ymax=1, xmin=-1, xmax=1) 
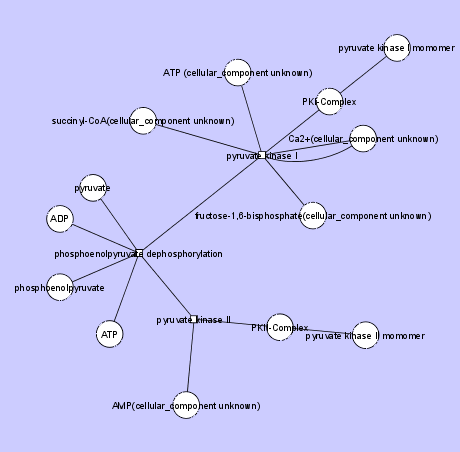
Here is the head: source target meansĬhoice of layout (fr or kk are both good for sparse graphs) l <- layout_with_fr(network_graph) Network_df is a ame with source, target and value. Netm <- get.adjacency(network_graph, sparse = F) Vertices = unique(as.character(network_df)) Network_graph <- graph_from_data_frame(d = network_df, directed = F) However, the network formatting (node colora etc) is lost! Is there a way to keep the formatting and transfer it to Cytoscape? My code library(igraph) I save my igraph object and use RC圓 R package (function createNetworkFromIgraph) to export it in cytoscape. I would like to import my network in Cytoscape in order to arrange the layout myself. I am using igraph R package to create and visualize a network of interest. 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
